Drop Plate Method for Counting Biofilm Cells
Student Version (go to Instructor Version)
Supplies Needed for Standard Method:
|
Quantity |
Description |
| biofilm | |
| 4 | sterile dilution tubes with caps (18 x 150 mm) |
| 1 | 1 ml automatic pipetting device |
| 4-5 | sterile 1 ml pipettes or pipette tips |
| As Necessary | Sterile buffered water |
| 4-5 | R2A agar plates per sample |
| 1 | 100 µl automatic pipetting device |
| 4-5 | 100 µl pipette tips |
| 1 | Sharpie marking pens |
| 1 | small ruler |
| 1 | Vortex mixer |
Instructions:
Serial Dilutions
- Pipette 1 ml of bacterial suspension (from the Harvesting exercise) into a dilution tube containing 9 ml of sterile buffered water.
- Recap tube and vortex the tube for approximately 8 seconds.
- Using a new tip, pipette 1 ml of vortexed dilution tube into a second dilution tube containing 9 ml of sterile buffered water.
- Repeat process until there are four serial dilutions of the original 9 ml sample. See Figure 1.
Supplies Needed for Alternate Method:
|
Quantity |
Description |
| As Necessary | biofilm |
| 4 | sterile dilution tubes with caps (18 x 150 mm) |
| 1 | 1 ml automatic pipetting device |
| 4-5 | sterile 1 ml pipettes or pipette tips |
| As Necessary | Sterile buffered water |
| 4-5 | R2A agar plates per sample |
| 1 | 100 µl automatic pipetting device |
| 4-5 | 100 µl pipette tips |
| 1 | Sharpie marking pens |
| 1 | small ruler |
| 1 | Vortex mixer |
Instructions:
Serial Dilutions
- Pipette 1 ml of bacterial suspension (from the Harvesting exercise) into a dilution tube containing 9 ml of sterile buffered water.
- Recap tube and vortex the tube for approximately 8 seconds.
- Using a new tip, pipette 1 ml of vortexed dilution tube into a second dilution tube containing 9 ml of sterile buffered water.
- epeat process until there are four serial dilutions of the original 9 ml sample. See Figure 2.
For both methods continue with these instructions
Drop Plating
- Label the bottom of the R2A agar plates by dividing them into fourths with a ruler and marking pen (see Figure 3). Each serial dilution of the sample will occupy one quadrant of each plate. Prepare plates in duplicate.
- Set the automatic pipetteman to pull up 100 µl of sample and expel 10 µl with each push of the button (slow setting).
- Vortex sample for approximately 8 seconds.
- Pick up sample with pippetteman. Expel sample in five evenly spaced 10 µl drops onto the quadrant of one of the petri plates that have been labeled for that particular dilution of the sample.
- Repeat this procedure with the duplicate plate. Note: If only a dedicated 10 ml pipette is available, 5 drops of 10 ml each should be placed on the plate.
- Vortex sample again for approximately 8 seconds.
- Pick up 100 µl of the next dilution and repeat steps 4-6.
- Repeat steps 4-6 for each tube in the dilution series.
- Let the drops soak into the media before turning plates over for incubation.
- Incubate overnight (17-20 hours) at 35-37˚C.
- Remove plates when colonies have developed and count the dilution which contains 3-30 colonies per 10 µl drop. Viable cell counts are expressed as colony forming units (CFU)/surface area.
Example Calculation
Calculate LOG10 CFU/cm2 according to the following
formula:
LOG (CFU/cm2) = LOG [(average CFU/drop volume)(dilution
counted)(volume scraped into/surface area*)] *scraped
slide
= LOG [(average of plate counts/ 0.01 ml)(10dilution)(10
ml/1.267cm2)]
For example, if the average units per drop was 16.6 and they
were counted in the third dilution, then the LOG (CFU/cm2) =
LOG [(16.6 / 0.01)(103)(10/1.267)] = 7.12
Illustrations:
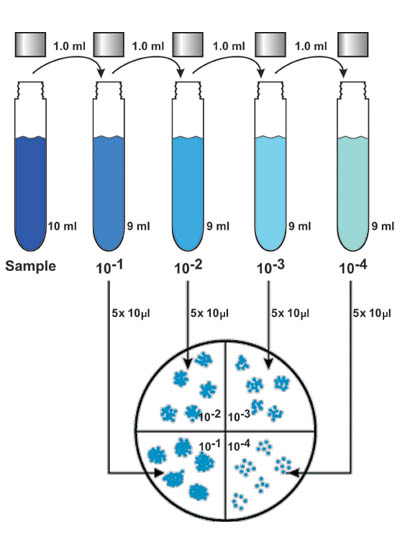
Figure 1. Dilution series followed by drop-plating technique for the Standard Method.
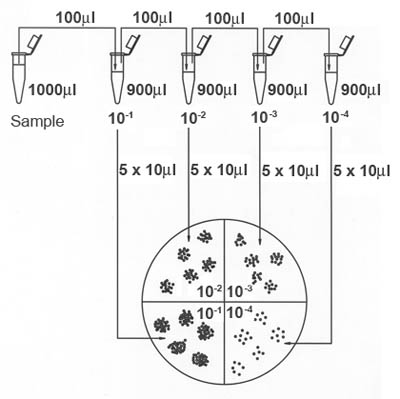
Figure 2. Dilution series followed by drop-plating technique for the Alternate Method.
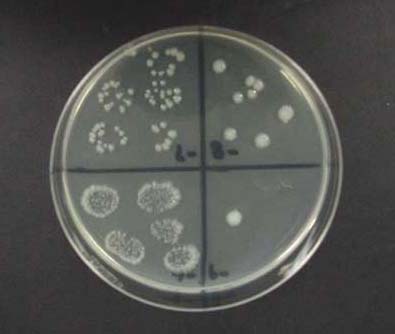
Figure 3. Example of a colonized agar plate, divided into four quadrants and containing five evenly spaced drops of diluent.
References:
How to optimize the drop plate method for enumerating
bacteria
Herigstad B, Hamilton M, Heersink J
J Microbiol Meth, 44(2): 121-129 (2001)
Measuring antimicrobial effects on biofilm bacteria: From
laboratory to field
Zelver N, Hamilton M, Pitts B, Goeres D, Walker D, Sturman P,
Heersink J
in R.J. Doyle, et al. (eds), Biofilms: Methods in Enzymology,
Academic Press, San Diego, CA, 1999, pp. 608-628.
Educational Program Curricula and Teaching Resources
Supported in part by the Waksman Foundation for MicrobiologyDeveloped in collaboration with Dr. John Lennox, Penn State University-Altoona
©1999-2006 Center for Biofilm Engineering, http://www.biofilm.montana.edu